Introduction
Create efficacy tables for clinical study reports: time-to-event analysis, response rates, subgroup analyses, and change from baseline endpoints.
Setup and Data Preparation
library(pharmhand)
library(dplyr)
library(tidyr)
library(ggplot2)
library(survival)
# Set flextable defaults for readable tables in light/dark mode
flextable::set_flextable_defaults(
font.color = "#000000",
background.color = "#FFFFFF"
)Use pharmaverseadam for example data:
library(pharmaverseadam)
# Load example datasets
adsl <- pharmaverseadam::adsl # Subject-level data
advs <- pharmaverseadam::advs # Vital signs data
adlb <- pharmaverseadam::adlb # Laboratory data
adtte <- pharmaverseadam::adtte_onco # Time-to-event data (oncology)
adrs <- pharmaverseadam::adrs_onco # Response data (oncology)
# Note: adtte_onco and adrs_onco use ARM as treatment variable (not TRT01P)Time-to-Event Analysis
create_tte_summary_table() provides survival analysis
with Kaplan-Meier estimates, hazard ratios, and landmark analyses.
Basic TTE
# Create TTE summary table
# Note: adtte_onco uses ARM for treatment groups
tte_table <- create_tte_summary_table(
data = adtte,
time_var = "AVAL",
event_var = "CNSR",
trt_var = "ARM", # Use ARM instead of TRT01P for oncology data
title = "Overall Survival Summary"
)
# Display the table
tte_table@flextableOverall Survival Summary | |||
|---|---|---|---|
Statistic |
Placebo |
Xanomeline High Dose |
Xanomeline Low Dose |
N |
174 |
169 |
169 |
Events n (%) |
5 (2.9) |
3 (1.8) |
2 (1.2) |
Median (95% CI) |
NE |
NE |
NE |
HR (95% CI) |
Reference |
0.84 (0.20, 3.53) |
0.56 (0.11, 2.92) |
p-value |
- |
0.809 |
0.492 |
FAS Population | |||
Time unit: months | |||
HR reference group: Placebo | |||
NE = Not Estimable | |||
TTE with Landmarks
# TTE table with landmark survival estimates
tte_landmark <- create_tte_summary_table(
data = adtte,
time_var = "AVAL",
event_var = "CNSR",
trt_var = "ARM",
landmarks = c(12, 24), # 12 and 24 month survival rates
time_unit = "months",
title = "Progression-Free Survival with Landmark Analysis"
)
# Display the table
tte_landmark@flextableProgression-Free Survival with Landmark Analysis | |||
|---|---|---|---|
Statistic |
Placebo |
Xanomeline High Dose |
Xanomeline Low Dose |
N |
174 |
169 |
169 |
Events n (%) |
5 (2.9) |
3 (1.8) |
2 (1.2) |
Median (95% CI) |
NE |
NE |
NE |
12-months Rate (95% CI) |
97.8 (94.8, 100.0) |
97.8 (94.8, 100.0) |
97.8 (94.8, 100.0) |
24-months Rate (95% CI) |
97.8 (94.8, 100.0) |
97.8 (94.8, 100.0) |
97.8 (94.8, 100.0) |
HR (95% CI) |
Reference |
0.84 (0.20, 3.53) |
0.56 (0.11, 2.92) |
p-value |
- |
0.809 |
0.492 |
FAS Population | |||
Time unit: months | |||
HR reference group: Placebo | |||
NE = Not Estimable | |||
Kaplan-Meier Plots
# Basic KM plot
km_plot <- create_km_plot(
data = adtte,
time_var = "AVAL",
event_var = "CNSR",
trt_var = "ARM",
title = "Overall Survival Kaplan-Meier Plot",
show_median = TRUE,
show_ci = TRUE,
xlab = "Days"
)
km_plot@plot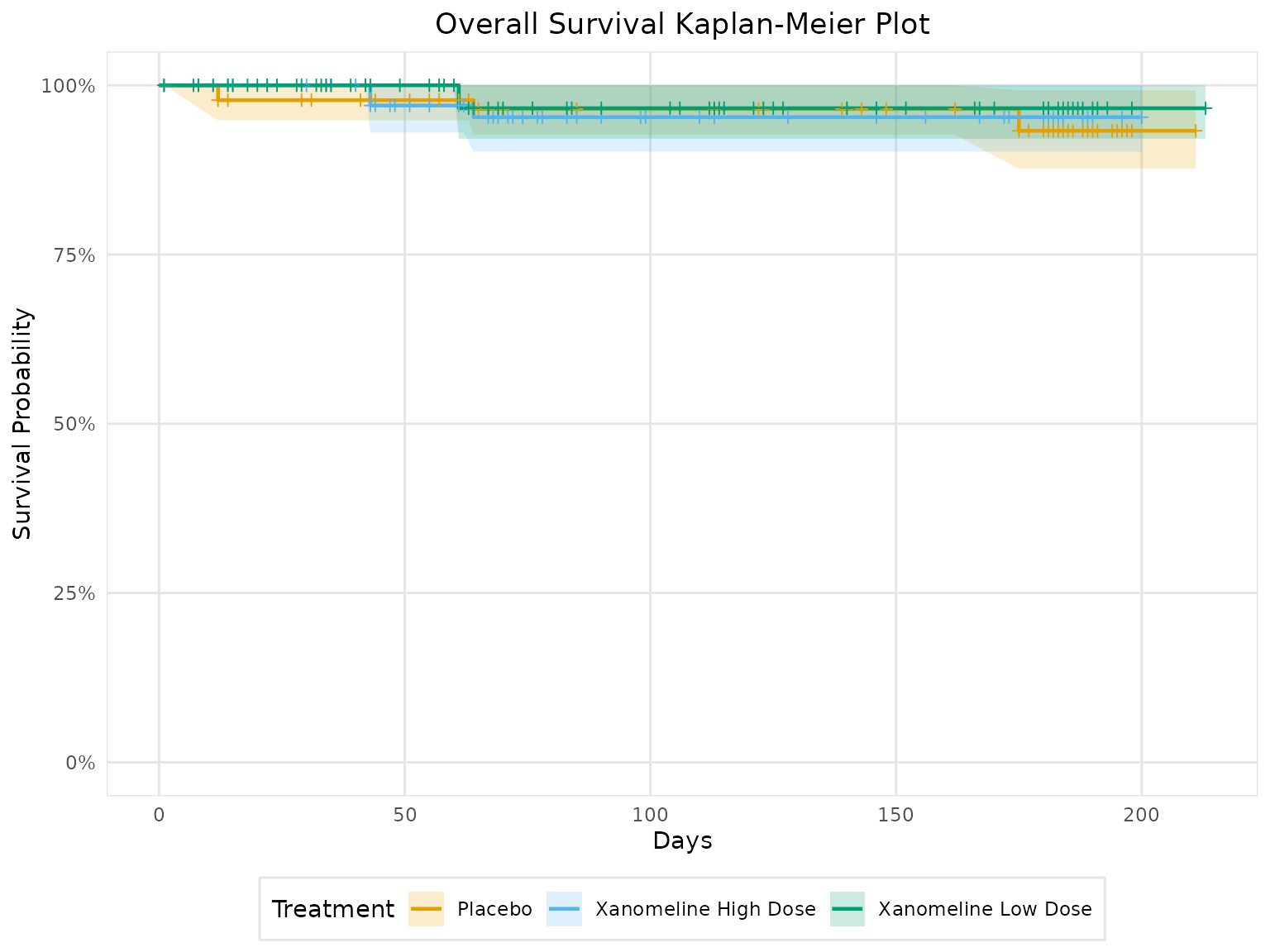
# Advanced KM plot with risk table
km_plot_advanced <- create_km_plot(
data = adtte,
time_var = "AVAL",
event_var = "CNSR",
trt_var = "ARM",
title = "Survival Analysis with Risk Table",
show_median = TRUE,
show_ci = TRUE,
xlab = "Days",
risk_table = TRUE,
landmarks = c(12, 24)
)
km_plot_advanced@plot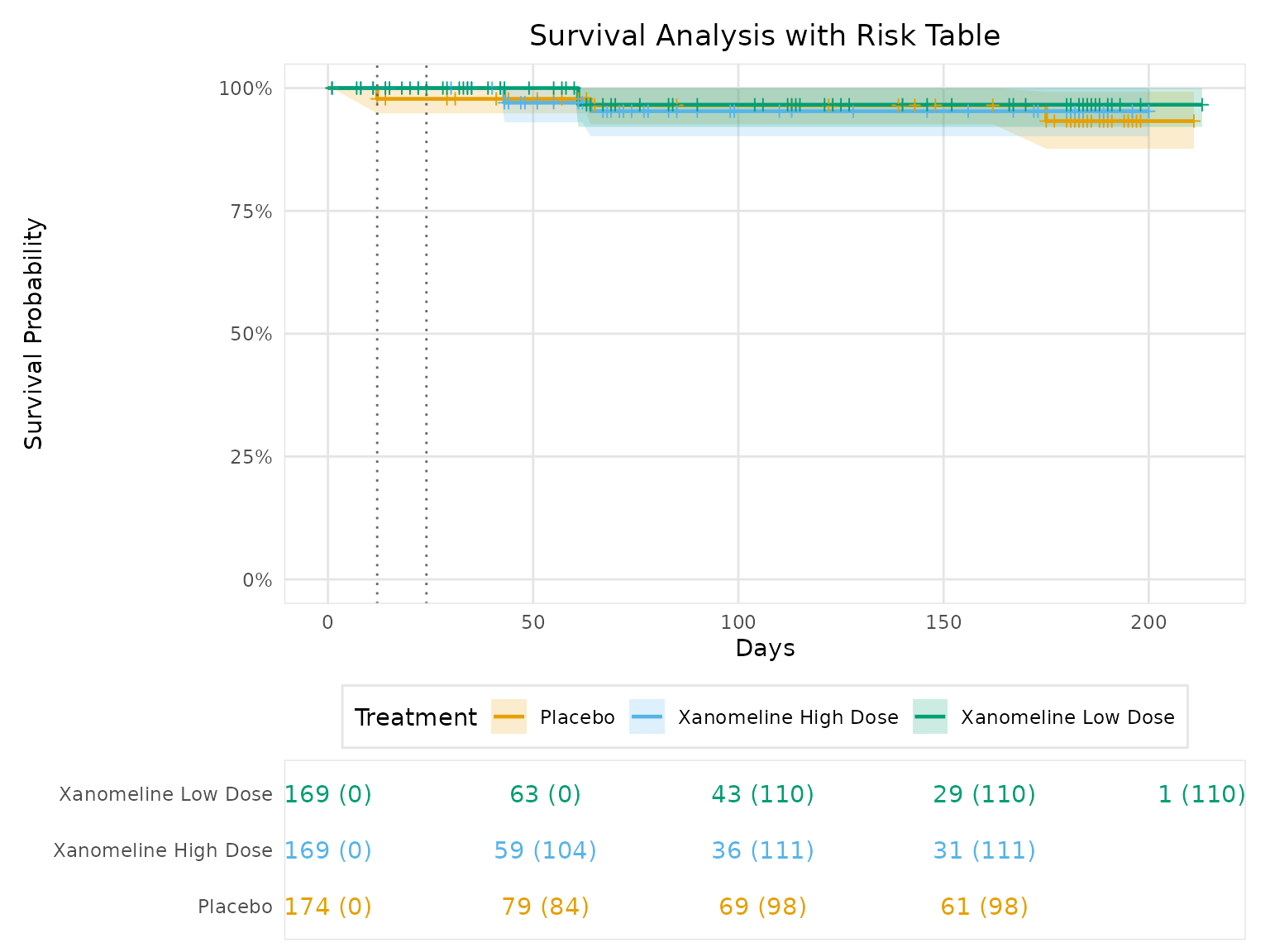
Proportional Hazards Diagnostics
Use Schoenfeld residual tests, log-log plots, and automatic warnings to validate Cox model assumptions across time-to-event analyses.
# Schoenfeld residual test with optional diagnostic plot
ph_results <- test_ph_assumption(
data = adtte,
time_var = "AVAL",
event_var = "CNSR",
trt_var = "ARM",
plot = TRUE
)
ph_results$results
ph_results$plot@plot
# Log-log survival plot for visual PH assessment
loglog_plot <- create_loglog_plot(
data = adtte,
time_var = "AVAL",
event_var = "CNSR",
trt_var = "ARM",
title = "Log-log Survival Plot"
)
loglog_plot@plot
# create_tte_summary_table() checks PH by default (check_ph = TRUE)Responder Analysis
create_responder_table() provides response summaries with multiple comparison options.
Basic Response
# Create response rate table for tumor response
# Note: adrs_onco uses ARM for treatment groups
response_table <- create_responder_table(
data = adrs,
response_var = "AVALC",
response_values = c("CR", "PR"), # CR + PR = Overall Response Rate
trt_var = "ARM",
title = "Best Overall Response Rate"
)
# Display the table
response_table@flextableBest Overall Response Rate | |||||
|---|---|---|---|---|---|
Treatment |
n/N |
Rate (%) |
95% CI |
OR (95% CI) |
p-value |
Placebo |
8/1042 |
0.8 |
(0.4, 1.5) |
Reference |
- |
Xanomeline High Dose |
3/1016 |
0.3 |
(0.1, 0.9) |
0.38 (0.10, 1.45) |
0.157 |
Xanomeline Low Dose |
4/1012 |
0.4 |
(0.2, 1.0) |
0.51 (0.15, 1.71) |
0.277 |
Screen Failure |
0/624 |
0.0 |
(0.0, 0.6) |
0.00 (0.00, Inf) |
0.989 |
Response defined as: CR, PR | |||||
95% CI calculated using wilson method | |||||
OR compared to Placebo | |||||
Response with Risk Ratio
# Response table using risk ratio instead of odds ratio
response_rr <- create_responder_table(
data = adrs,
response_var = "AVALC",
response_values = c("CR", "PR"),
trt_var = "ARM",
comparison_type = "RR", # Risk Ratio
ci_method = "wilson", # Wilson confidence interval
title = "Response Rate Summary (Risk Ratio)"
)
# Display the table
response_rr@flextableResponse Rate Summary (Risk Ratio) | |||||
|---|---|---|---|---|---|
Treatment |
n/N |
Rate (%) |
95% CI |
RR (95% CI) |
p-value |
Placebo |
8/1042 |
0.8 |
(0.4, 1.5) |
Reference |
- |
Xanomeline High Dose |
3/1016 |
0.3 |
(0.1, 0.9) |
0.38 (0.10, 1.45) |
0.226 |
Xanomeline Low Dose |
4/1012 |
0.4 |
(0.2, 1.0) |
0.51 (0.16, 1.70) |
0.387 |
Screen Failure |
0/624 |
0.0 |
(0.0, 0.6) |
0.10 (0.01, 1.70) |
0.029 |
Response defined as: CR, PR | |||||
95% CI calculated using wilson method | |||||
RR compared to Placebo | |||||
Response with Risk Difference
# Response table with risk difference
response_rd <- create_responder_table(
data = adrs,
response_var = "AVALC",
response_values = c("CR", "PR"),
trt_var = "ARM",
comparison_type = "RD", # Risk Difference
title = "Response Rate Difference"
)
# Display the table
response_rd@flextableResponse Rate Difference | |||||
|---|---|---|---|---|---|
Treatment |
n/N |
Rate (%) |
95% CI |
RD (95% CI) |
p-value |
Placebo |
8/1042 |
0.8 |
(0.4, 1.5) |
Reference |
- |
Xanomeline High Dose |
3/1016 |
0.3 |
(0.1, 0.9) |
-0.5 (-1.1, 0.2) |
0.139 |
Xanomeline Low Dose |
4/1012 |
0.4 |
(0.2, 1.0) |
-0.4 (-1.0, 0.3) |
0.266 |
Screen Failure |
0/624 |
0.0 |
(0.0, 0.6) |
-0.8 (-1.3, -0.2) |
0.005 |
Response defined as: CR, PR | |||||
95% CI calculated using wilson method | |||||
RD compared to Placebo | |||||
Subgroup Analysis
Subgroup analyses show treatment effects across patient populations.
Subgroup Table
# Create subgroup analysis table
subgroup_table <- create_subgroup_table(
data = adtte,
subgroups = list(
SEX = "Sex"
),
endpoint_type = "tte", # Time-to-event endpoint
trt_var = "ARM",
title = "Subgroup Analysis - Overall Survival"
)
# Display the table
subgroup_table@flextableSubgroup Analysis - Overall Survival | |||||
|---|---|---|---|---|---|
Subgroup |
Category |
n (Placebo) |
n (Xanomeline High Dose) |
HR (95% CI) |
Interaction p |
Overall |
174 |
338 |
0.84 (0.20, 3.53) |
||
Sex |
F |
107 |
181 |
1.49 (0.21, 10.61) |
0.249 |
Sex |
M |
67 |
157 |
0.47 (0.05, 4.62) |
|
HR = Hazard Ratio | |||||
Reference group: Placebo | |||||
Interaction p-value from likelihood ratio test | |||||
NE = Not Estimable | |||||
Forest Plot
# Create forest plot for subgroup analysis
forest_plot <- create_forest_plot(
data = adtte,
subgroups = list(
SEX = "Sex"
),
endpoint_type = "tte",
trt_var = "ARM",
title = "Treatment Effect by Subgroup",
show_interaction = TRUE
)
forest_plot@plot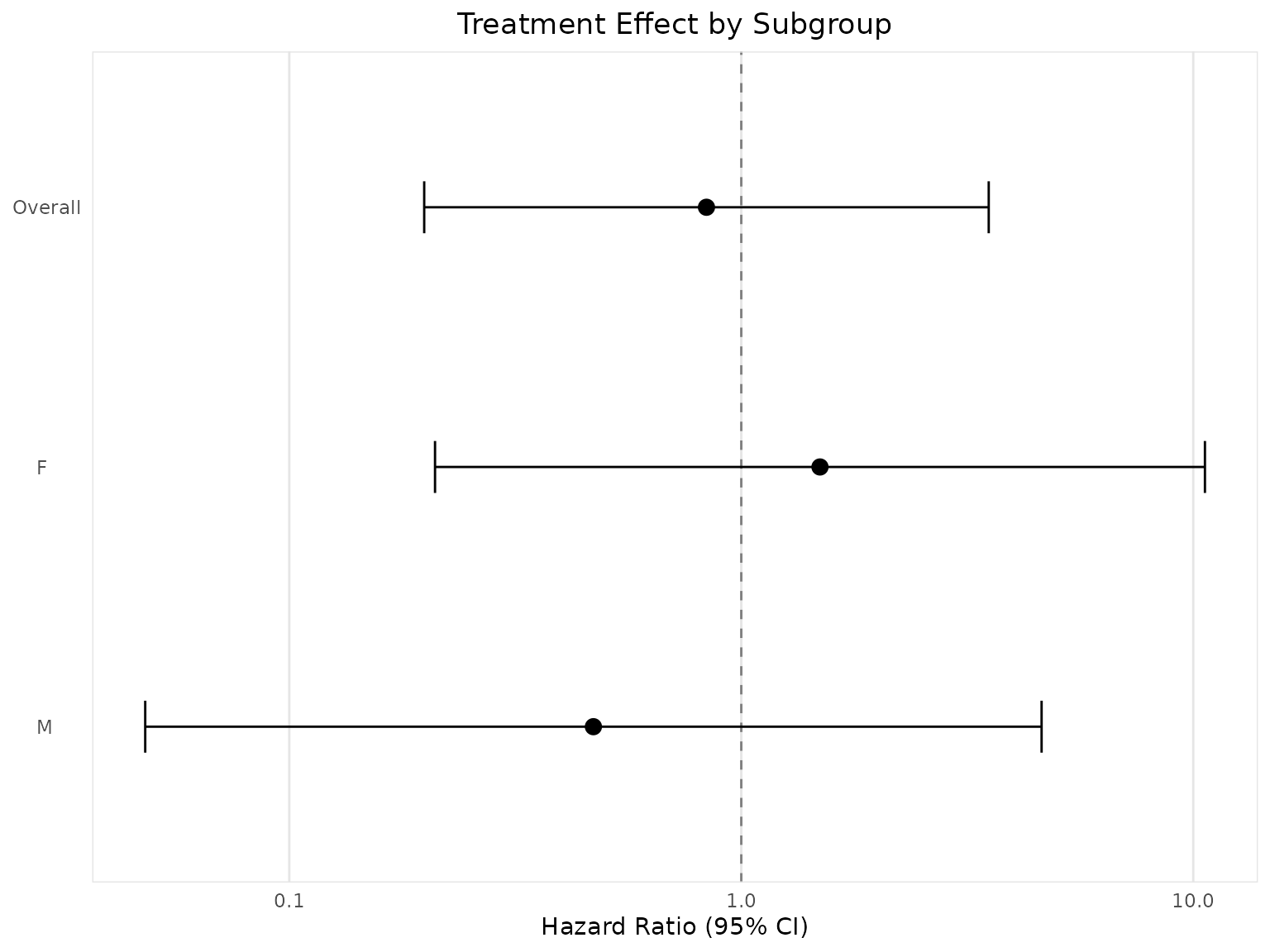
Binary Endpoint
# Subgroup analysis for binary endpoint
subgroup_binary <- create_subgroup_table(
data = adrs,
subgroups = list(
SEX = "Sex"
),
endpoint_type = "binary", # Binary endpoint
response_var = "AVALC",
response_values = c("CR", "PR"),
trt_var = "ARM",
title = "Response Rate by Subgroup"
)
# Display the table
subgroup_binary@flextableResponse Rate by Subgroup | |||||
|---|---|---|---|---|---|
Subgroup |
Category |
n (Placebo) |
n (Screen Failure) |
OR (95% CI) |
Interaction p |
Overall |
1042 |
2652 |
0.00 (0.00, Inf) |
||
Sex |
F |
639 |
1517 |
0.00 (0.00, Inf) |
0.009 |
Sex |
M |
403 |
1135 |
0.00 (0.00, Inf) |
|
OR = Odds Ratio | |||||
Reference group: Placebo | |||||
Interaction p-value from likelihood ratio test | |||||
NE = Not Estimable | |||||
Change from Baseline Analysis
create_cfb_summary_table() analyzes change from baseline
values for continuous endpoints.
Vital Signs
# Create change from baseline table for vital signs
cfb_table <- create_cfb_summary_table(
data = advs,
params = c("SYSBP", "DIABP", "PULSE"),
visit = "End of Treatment",
title = "Change from Baseline in Vital Signs"
)
# Display the table
cfb_table@flextableChange from Baseline in Vital Signs | ||||||
|---|---|---|---|---|---|---|
Parameter |
Placebo n |
Xanomeline High Dose n |
Xanomeline Low Dose n |
Placebo Mean (SD) |
Xanomeline High Dose Mean (SD) |
Xanomeline Low Dose Mean (SD) |
Diastolic Blood Pressure (mmHg) |
222 |
168 |
177 |
-3 (9) |
-2.74 (11.34) |
0.11 (10.12) |
Pulse Rate (beats/min) |
222 |
168 |
177 |
1.63 (10.7) |
1.2 (8.93) |
4.29 (10.39) |
Systolic Blood Pressure (mmHg) |
222 |
168 |
177 |
-3.56 (17.82) |
-6.74 (14.96) |
-3.7 (14.06) |
FAS Population | ||||||
Mean (SD) presented for change from baseline | ||||||
Custom Parameters
# Change from baseline for specific parameters
cfb_custom <- create_cfb_summary_table(
data = advs,
params = c("SYSBP"), # Only systolic blood pressure
visit = "Week 8",
title = "Systolic Blood Pressure Change from Baseline at Week 8"
)
# Display the table
cfb_custom@flextableSystolic Blood Pressure Change from Baseline at Week 8 | ||||||
|---|---|---|---|---|---|---|
Parameter |
Placebo n |
Xanomeline High Dose n |
Xanomeline Low Dose n |
Placebo Mean (SD) |
Xanomeline High Dose Mean (SD) |
Xanomeline Low Dose Mean (SD) |
Systolic Blood Pressure (mmHg) |
219 |
168 |
180 |
0.05 (15.09) |
-3.64 (15.93) |
-1.17 (17.25) |
FAS Population | ||||||
Mean (SD) presented for change from baseline | ||||||
Primary Endpoint Tables
create_primary_endpoint_table() creates summary tables for primary and secondary endpoints.
Primary Endpoint
# Create primary endpoint summary table
primary_table <- create_primary_endpoint_table(
data = advs,
paramcd = "SYSBP",
visit = "End of Treatment",
title = "Primary Endpoint Summary: Systolic Blood Pressure"
)
# Display the table
primary_table@flextablePrimary Endpoint Summary: Systolic Blood Pressure | |||
|---|---|---|---|
Statistic |
Placebo |
Xanomeline High Dose |
Xanomeline Low Dose |
n |
222 |
168 |
177 |
Mean (SD) |
132.7 (15.44) |
132.3 (15.62) |
133 (17.08) |
Median |
131 |
131 |
130 |
Min, Max |
78, 172 |
100, 177 |
92, 178 |
FAS Population | |||
Parameter: SYSBP at End of Treatment | |||
Secondary Endpoint
# Secondary endpoint analysis
secondary_table <- create_primary_endpoint_table(
data = advs,
paramcd = "DIABP",
visit = "Week 8",
title = "Secondary Endpoint: Diastolic Blood Pressure at Week 8"
)
# Display the table
secondary_table@flextableSecondary Endpoint: Diastolic Blood Pressure at Week 8 | |||
|---|---|---|---|
Statistic |
Placebo |
Xanomeline High Dose |
Xanomeline Low Dose |
n |
292 |
224 |
240 |
Mean (SD) |
75.2 (9.12) |
77.4 (9.07) |
75.4 (10.59) |
Median |
76 |
78.3 |
74 |
Min, Max |
49, 101 |
54, 98 |
52, 100 |
FAS Population | |||
Parameter: DIABP at Week 8 | |||
Laboratory Analysis
Analyze laboratory parameters with shift tables and summary statistics.
Lab Summary
Click to expand: Laboratory Parameters Summary
# Create laboratory parameters summary
lab_summary <- create_lab_summary_table(
data = adlb,
params = c("HGB", "WBC", "PLAT", "ALT", "AST"),
visit = "Week 24",
title = "Laboratory Parameters Summary at Week 24"
)
# Display the table
lab_summary@flextableLaboratory Parameters Summary at Week 24 | ||||||
|---|---|---|---|---|---|---|
Parameter |
Placebo n |
Xanomeline High Dose n |
Xanomeline Low Dose n |
Placebo Mean (SD) |
Xanomeline High Dose Mean (SD) |
Xanomeline Low Dose Mean (SD) |
Alanine Aminotransferase (U/L) |
57 |
30 |
26 |
17.86 (15.61) |
20.97 (8.7) |
18.19 (9.17) |
Aspartate Aminotransferase (U/L) |
57 |
30 |
26 |
25.19 (21.02) |
24.43 (7.29) |
22.42 (10.78) |
Hemoglobin (mmol/L) |
58 |
30 |
25 |
8.32 (0.82) |
8.88 (0.85) |
8.38 (0.69) |
Platelet (10^9/L) |
57 |
29 |
24 |
238.75 (51.89) |
238.34 (67.53) |
249.71 (63.44) |
Leukocytes (10^9/L) |
58 |
30 |
25 |
6.67 (1.77) |
6.74 (1.8) |
6.29 (1.84) |
Safety Population | ||||||
Mean (SD) presented | ||||||
Lab Shift
Click to expand: Lab Shift Table
# Create laboratory shift table for liver function test
shift_table <- create_lab_shift_table(
data = adlb,
paramcd = "ALT",
visit = "Week 24",
title = "ALT Shift from Baseline to Week 24"
)
# Display the table
shift_table@flextableALT Shift from Baseline to Week 24 | ||||
|---|---|---|---|---|
Planned Treatment for Period 01 |
Baseline Reference Range Indicator |
NORMAL |
HIGH |
LOW |
Placebo |
NORMAL |
51 |
2 |
1 |
Xanomeline High Dose |
NORMAL |
28 |
0 |
0 |
Xanomeline Low Dose |
NORMAL |
24 |
2 |
0 |
Xanomeline High Dose |
HIGH |
2 |
0 |
0 |
Placebo |
HIGH |
3 |
0 |
0 |
Safety Population | ||||
Shift from baseline normal range indicator | ||||
Vital Signs by Visit
create_vs_by_visit_table() tracks vital signs across study visits.
Click to expand: Vital Signs by Visit Table
# Vital signs by visit table
vs_visit_table <- create_vs_by_visit_table(
data = advs,
paramcd = "SYSBP",
visits = c("Baseline", "Week 2", "Week 4", "Week 8", "End of Treatment"),
title = "Systolic Blood Pressure by Study Visit"
)
# Display the table
vs_visit_table@flextableSystolic Blood Pressure by Study Visit | |||
|---|---|---|---|
Visit |
Placebo |
Xanomeline High Dose |
Xanomeline Low Dose |
Baseline |
340 / 136.8 (17.57) |
336 / 138.8 (18.52) |
336 / 136.9 (17.27) |
Week 2 |
336 / 133.8 (17.54) |
325 / 132.3 (16.37) |
335 / 135.2 (17.42) |
Week 4 |
328 / 133.9 (17.37) |
292 / 132.2 (16.14) |
288 / 135.4 (17.79) |
Week 8 |
292 / 136.3 (17.02) |
224 / 135.1 (15.54) |
240 / 134.9 (17.84) |
End of Treatment |
222 / 132.7 (15.44) |
168 / 132.3 (15.62) |
177 / 133 (17.08) |
FAS Population | |||
Format: n / Mean (SD) | |||
Customization
Functions include options for confidence intervals, formatting, and custom parameters.
Formatting
# Custom formatting options
custom_table <- create_tte_summary_table(
data = adtte,
time_var = "AVAL",
event_var = "CNSR",
trt_var = "ARM",
conf_level = 0.95,
time_unit = "months",
title = "Custom Formatted TTE Table",
autofit = TRUE # Auto-adjust column widths
)
# Display the table
custom_table@flextableCustom Formatted TTE Table | |||
|---|---|---|---|
Statistic |
Placebo |
Xanomeline High Dose |
Xanomeline Low Dose |
N |
174 |
169 |
169 |
Events n (%) |
5 (2.9) |
3 (1.8) |
2 (1.2) |
Median (95% CI) |
NE |
NE |
NE |
HR (95% CI) |
Reference |
0.84 (0.20, 3.53) |
0.56 (0.11, 2.92) |
p-value |
- |
0.809 |
0.492 |
FAS Population | |||
Time unit: months | |||
HR reference group: Placebo | |||
NE = Not Estimable | |||
CI Methods
# Different CI methods for response rates
wilson_table <- create_responder_table(
data = adrs,
response_var = "AVALC",
response_values = c("CR", "PR"),
trt_var = "ARM",
ci_method = "wilson", # Wilson score interval (recommended)
title = "Response Rate - Wilson CI"
)
exact_table <- create_responder_table(
data = adrs,
response_var = "AVALC",
response_values = c("CR", "PR"),
trt_var = "ARM",
ci_method = "exact", # Clopper-Pearson exact
title = "Response Rate - Exact CI"
)
# Display the tables
wilson_table@flextableResponse Rate - Wilson CI | |||||
|---|---|---|---|---|---|
Treatment |
n/N |
Rate (%) |
95% CI |
OR (95% CI) |
p-value |
Placebo |
8/1042 |
0.8 |
(0.4, 1.5) |
Reference |
- |
Xanomeline High Dose |
3/1016 |
0.3 |
(0.1, 0.9) |
0.38 (0.10, 1.45) |
0.157 |
Xanomeline Low Dose |
4/1012 |
0.4 |
(0.2, 1.0) |
0.51 (0.15, 1.71) |
0.277 |
Screen Failure |
0/624 |
0.0 |
(0.0, 0.6) |
0.00 (0.00, Inf) |
0.989 |
Response defined as: CR, PR | |||||
95% CI calculated using wilson method | |||||
OR compared to Placebo | |||||
exact_table@flextableResponse Rate - Exact CI | |||||
|---|---|---|---|---|---|
Treatment |
n/N |
Rate (%) |
95% CI |
OR (95% CI) |
p-value |
Placebo |
8/1042 |
0.8 |
(0.3, 1.5) |
Reference |
- |
Xanomeline High Dose |
3/1016 |
0.3 |
(0.1, 0.9) |
0.38 (0.10, 1.45) |
0.157 |
Xanomeline Low Dose |
4/1012 |
0.4 |
(0.1, 1.0) |
0.51 (0.15, 1.71) |
0.277 |
Screen Failure |
0/624 |
0.0 |
(0.0, 0.6) |
0.00 (0.00, Inf) |
0.989 |
Response defined as: CR, PR | |||||
95% CI calculated using exact method | |||||
OR compared to Placebo | |||||
Complete Report
Combine multiple analyses into a report:
generate_efficacy_report <- function(output_path = "Efficacy_Report.docx") {
# Section 1: Primary Endpoint Summary
primary_content <- create_primary_endpoint_table(
data = advs,
paramcd = "SYSBP",
visit = "End of Treatment",
title = "Table 3.1: Primary Endpoint Summary (Systolic BP)"
)
# Section 2: Change from Baseline
cfb_content <- create_cfb_summary_table(
data = advs,
params = c("SYSBP", "DIABP", "PULSE"),
visit = "End of Treatment",
title = "Table 3.2: Change from Baseline Summary"
)
# Section 3: Vital Signs by Visit
vs_content <- create_vs_by_visit_table(
data = advs,
paramcd = "SYSBP",
title = "Table 3.3: Vital Signs by Visit"
)
# Section 4: Laboratory Parameters
lab_content <- create_lab_summary_table(
data = adlb,
title = "Table 3.4: Laboratory Parameters Summary"
)
# Section 5: Laboratory Shift Table
shift_content <- create_lab_shift_table(
data = adlb,
paramcd = "ALT",
title = "Table 3.5: ALT Shift from Baseline"
)
# Section 6: Subgroup Analysis
subgroup_content <- create_subgroup_analysis_table(
adsl = adsl,
data = advs,
title = "Table 3.6: Subgroup Analysis by Age Group"
)
# Create report sections
sections <- list(
ReportSection(
title = "Primary Endpoint Analysis",
section_type = "efficacy",
content = list(primary_content)
),
ReportSection(
title = "Change from Baseline Analysis",
section_type = "efficacy",
content = list(cfb_content)
),
ReportSection(
title = "Vital Signs by Study Visit",
section_type = "efficacy",
content = list(vs_content)
),
ReportSection(
title = "Laboratory Parameters",
section_type = "efficacy",
content = list(lab_content)
),
ReportSection(
title = "Laboratory Shift Analysis",
section_type = "efficacy",
content = list(shift_content)
),
ReportSection(
title = "Subgroup Analyses",
section_type = "efficacy",
content = list(subgroup_content)
)
)
# Create and save report
report <- ClinicalReport(
study_id = "CDISCPILOT01",
study_title = "CDISC Pilot Study - Efficacy Report",
sections = sections,
metadata = list(
generated_at = Sys.time(),
package_version = as.character(packageVersion("pharmhand")),
data_source = "pharmaverseadam",
report_type = "efficacy"
)
)
# Generate Word document
generate_word(report, path = output_path)
cat("Efficacy report generated:", output_path, "\n")
invisible(report)
}
# Generate the report
generate_efficacy_report()